Background:Mantle cell lymphoma (MCL) is the secondary common B cell lymphoma subtype that comprises 6 to 8% of non-Hodgkin's lymphoma, and is closely related to the poor clinical outcomes. Previous studies have focussed on the mechanisms mediating ibrutinib resistance, the first-in-class oral covalent inhibitor of Bruton's tyrosine kinase (BTK). The aims of the this study is to identify key genes related to the FGFR1 Knockdown in Mantle cell lymphoma cell line (Z-138) that has been proved to play a vital role in MCL progression.
Methods:GSE138127 mRNA microarray datasets from Gene Expression Omnibus (GEO) were analysed to obtain differentially expressed genes (DEGs) between vector control and the Knockdown of FGFR1 charactered by ibrutinib resistance. The GO and GSEA were carried out by WEB-based GEne SeT AnaLysis Toolkit (WebGestalt) to do the functional enrichment analysis. The network analysis of protein-protein interactions (PPIs), TF-gene interaction were carried out by Network Analyst 3.0 to identify hub genes. And the main hub gene was probed using Kyoto Encyclopedia of Genes and Genomes(KEGG) pathway analysis.
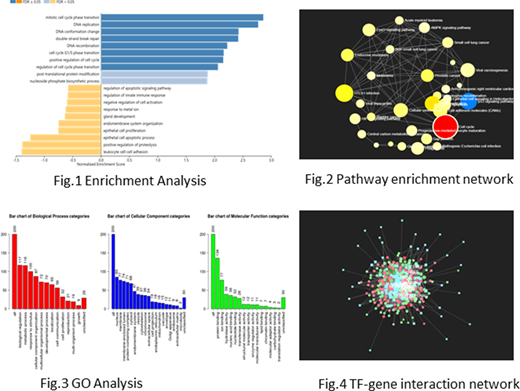
Results:In total, 175 DEGs were obtained, of which 87 and 88 were up- and down-regulated, respectively. Three hub genes (CDK1, CCND1) were identified and associated to cell cycle and DNA replication. FOXC1 were identified as the potential Transcription factors in the biological process.
Conclusion:CDK1, CCND1 may affect the cell cycle regulated by FOXC1, and represent the new candidate molecular markers of the occurrence of ibrutinib resistance.
Keywords: Mantle cell lymphoma (MCL), ibrutinib resistance, Network Analyst, Microarray, Protein-protein interactions, Molecular markers
No relevant conflicts of interest to declare.
Author notes
Asterisk with author names denotes non-ASH members.

This feature is available to Subscribers Only
Sign In or Create an Account Close Modal